Feedback
CrateDB with R¶
This integration document details how to create a Machine Learning pipeline using R and CrateDB.
Abstract¶
Statistical analysis and visualization on huge datasets is a common task many data scientists face in their day-to-day life. One common tool for doing this is R - a programming language and software environment geared towards statistical computing. CrateDB can be paired with R to provide a source of large, real-time machine-data for use in statistical analysis, visualization and machine learning.
Similarly, CrateDB can also be used by R as a store of the results of these statistical computations.
This can be accomplished with the RPostgreSQL library.
Implementation¶
Set Up¶
For this implementation, we will be using the classic iris classification problem.
We have details about three species of iris: iris setosa, iris versicolor, and iris virginica. These three species can be differentiated by the width and length of their sepals and of their petals.
We have the details about these irises stored in a CrateDB table, doc.iris.
We also have details about irises that we do not know the species of stored in
a CrateDB table, doc.unclassified_iris.
Using R, we want to:
Retrieve data from a CrateDB instance.
Perform some data visualization so that we can see the differences between the three iris species.
Build a machine learning model from this data that will predict an iris species given the four features (sepal and petal width and length) we have measured.
Retrieve our unclassified iris data, enrich the data with a prediction from our model, and insert the result into our iris table.
Prerequisites¶
- R (optionally with a third party tool like RStudio)
The RPostgreSQL library
The caret library
To install these libraries within R or RStudio, we can run:
> install.packages("RPostgreSQL")
> install.packages("caret")
CrateDB¶
First, we need to create a table to hold our training data, as well as our unclassified irises:
CREATE TABLE iris
(
sepal_length DOUBLE,
sepal_width DOUBLE,
petal_length DOUBLE,
petal_width DOUBLE,
species TEXT
);
CREATE TABLE unclassified_iris
(
sepal_length DOUBLE,
sepal_width DOUBLE,
petal_length DOUBLE,
petal_width DOUBLE
);
Once these tables have been created, we can import the iris data:
COPY iris FROM 'file:///path/to/iris.csv';
COPY unclassified_iris from 'file:///path/to/unclassified_iris.csv';
We can verify that the data has been successfully imported like so:
SELECT COUNT(*) FROM iris;
+----------+
| count(*) |
+----------+
| 150 |
+----------+
SELECT 1 row in set (0.130 sec)
Examining The Data¶
With our data in CrateDB, we can now load it into R or RStudio. Within
R, we should first import our data. We do this by loading the RPostgreSQL
library, connecting to a CrateDB database and loading the dataset:
# Loading the DBI library.
> library(DBI)
# Opening a connection, where dbname is the name of our schema, host is the address
# of the CrateDB instance, port is the CrateDB Psql port, and user is the default
# CrateDB user (in this case "crate").
> con <- dbConnect(RPostgres::Postgres(),
dbname = "doc",
host = "localhost",
port = 5432,
user = "crate")
# Loading the iris dataset from CrateDB
> iris_dataset <- dbGetQuery(con, "SELECT * FROM doc.iris")
We can then validate that we have loaded the dataset correctly by looking at the dimensions and the summary of the dataset:
# Getting the dimensions of the dataset.
> dim(iris_dataset)
[1] 150 5
# This indicates it contains 150 instances across 5 attributes. We can check
# the types of those attributes as well.
> sapply(iris_dataset, class)
sepal_length sepal_width petal_length petal_width species
"numeric" "numeric" "numeric" "numeric" "character"
# Finally, we can look at the the statistical summary of our dataset.
> summary(iris_dataset)
sepal_length sepal_width petal_length petal_width
Min. :4.300 Min. :2.000 Min. :1.000 Min. :0.100
1st Qu.:5.100 1st Qu.:2.800 1st Qu.:1.600 1st Qu.:0.300
Median :5.800 Median :3.000 Median :4.350 Median :1.300
Mean :5.843 Mean :3.054 Mean :3.759 Mean :1.199
3rd Qu.:6.400 3rd Qu.:3.300 3rd Qu.:5.100 3rd Qu.:1.800
Max. :7.900 Max. :4.400 Max. :6.900 Max. :2.500
species
Length:150
Class :character
Mode :character
We can now visualize our data. For example, visualizing a boxplot of iris properties by iris species might give us an insight about the distribution of these properties across each species.
# Boxplot of iris features by each species
> par(mfrow=c(2,2))
> boxplot(sepal_length~species, data=iris_dataset, main="Sepal Length by Species")
> boxplot(sepal_width~species, data=iris_dataset, main="Sepal Width by Species")
> boxplot(petal_length~species, data=iris_dataset, main="Petal Length by Species")
> boxplot(petal_width~species, data=iris_dataset, main="Petal Width by Species")
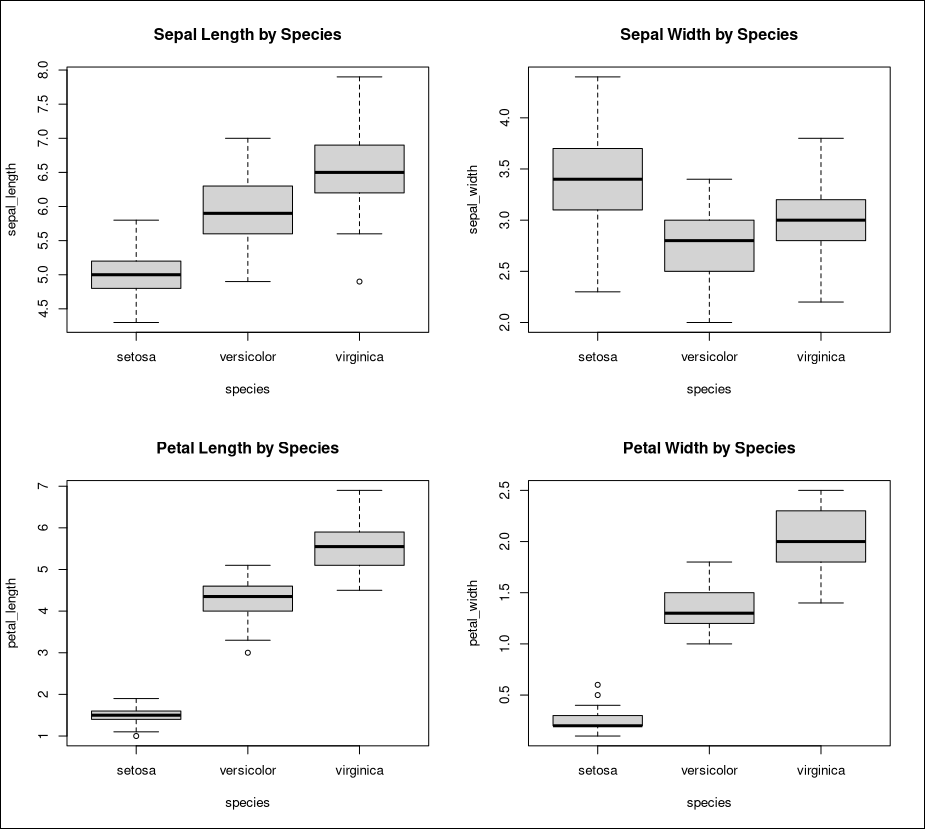
As we can see, the lengths and widths of sepals and petals are very good indicators of iris species, with little overlap between them.
Training A Model¶
Now that we have loaded our data and can visualize it to get a better idea of what it contains, we can create a machine learning model to predict a species of iris given sepal length/width and petal length/width.
For this, we will use Linear Discriminant Analysis (LDA), a dimensionality reduction technique often used in pattern classification, as is our goal here.
First, we will split our dataset into a set that contains 80% of the elements, for training, and 20% of the elements, for use in validating our model:
# Creating a partition that contains 80% of the dataset
validation_idx <- createDataPartition(iris_dataset$species, p=0.80, list=FALSE)
# Creating a dataset that contains 20% of the initial dataset for validation
> validation_dataset <- iris_dataset[-validation_idx,]
# Using the remaining 80% of the dataset for training.
training_dataset <- iris_dataset[validation_idx,]
# We can examine the dimensions of our datasets to verify the results.
> dim(training_dataset)
[1] 120 5
> dim(validation_dataset)
[1] 30 5
We now have a dataset suitable for training and a dataset suitable for validation. We can train an LDA model on this data, to predict the species based on a flower’s features.
# Importing the caret library
> library(caret)
# Training an LDA model, using the accuracy of the model to judge its effectiveness,
# and controlling the training using a 15-fold cross-validation.
> lda_model <- train(species~.,
data=training_dataset,
method="lda",
metric="Accuracy",
trControl=trainControl(method="cv", number=15))
Once this is trained, we can retrieve a summary of our model:
> print(lda_model)
Linear Discriminant Analysis
120 samples
4 predictor
3 classes: 'setosa', 'versicolor', 'virginica'
No pre-processing
Resampling: Cross-Validated (15 fold)
Summary of sample sizes: 113, 111, 111, 112, 113, 114, ...
Resampling results:
Accuracy Kappa
0.9916667 0.9873016
Our final model has an accuracy of 99.1%, which is pretty good. We can test our model on our verification dataset, and summarize the results in a confusion matrix:
# Create some predictions from our validation dataset.
> predictions <- predict(lda_model, validation_dataset)
# Comparing our predictions against the actual dataset via a confusion matrix.
> confusionMatrix(table(predictions, validation_dataset$species))
Confusion Matrix and Statistics
predictions setosa versicolor virginica
setosa 10 0 0
versicolor 0 9 1
virginica 0 1 9
Overall Statistics
Accuracy : 0.9333
95% CI : (0.7793, 0.9918)
No Information Rate : 0.3333
P-Value [Acc > NIR] : 8.747e-12
Our basic model looks to have predicted the results of our validation dataset with 93% accuracy - it has predicted all the setosa irises correctly, but misclassified a versicolor as a virginica and vice versa. We could improve this by trying out other models, by tweaking our model, or by training on a larger dataset.
Enriching Data¶
Now that we have a model we are happy with, we can use this model to enrich unclassified iris flowers data.
Within CrateDB we have a table, doc.unclassified_iris, that contains
irises without their classifications, which we can pull into R.
# Retrieving the dataset.
unclassified_dataset <- dbGetQuery(con, "SELECT * FROM doc.unclassified_iris")
> unclassified_dataset
sepal_length sepal_width petal_length petal_width
1 5.4 3.9 1.3 0.4
2 4.9 2.4 3.3 1.0
3 6.6 2.9 4.6 1.3
4 6.5 3.0 5.5 1.8
5 5.1 3.5 1.4 0.3
6 7.7 3.8 6.7 2.2
7 5.7 4.4 1.5 0.4
8 5.2 2.7 3.9 1.4
9 6.3 3.3 4.7 1.6
10 7.7 2.6 6.9 2.3
11 6.0 2.2 5.0 1.5
Using our LDA model, we can predict what species each of these are, and enrich our unclassified dataset with the species predictions:
# Creating a copy of our unclassified set.
> classified_dataset <- unclassified_dataset
# Enriching the dataset with a species prediction.
> classified_dataset$species <- predict(lda_model, unclassified_dataset)
# Our resulting dataset.
> classified_dataset
sepal_length sepal_width petal_length petal_width species
1 5.4 3.9 1.3 0.4 setosa
2 4.9 2.4 3.3 1.0 versicolor
3 6.6 2.9 4.6 1.3 versicolor
4 6.5 3.0 5.5 1.8 virginica
5 5.1 3.5 1.4 0.3 setosa
6 7.7 3.8 6.7 2.2 virginica
7 5.7 4.4 1.5 0.4 setosa
8 5.2 2.7 3.9 1.4 versicolor
9 6.3 3.3 4.7 1.6 versicolor
10 7.7 2.6 6.9 2.3 virginica
11 6.0 2.2 5.0 1.5 virginica
And finally insert the newly classified iris flowers into our iris table:
> query <- "INSERT INTO iris (sepal_length, sepal_width, petal_length, petal_width, species) values ( %s, %s, %s, %s, '%s')"
> for (i in 1:(dim(unclassified_dataset)[1]) ) {
+ q <- sprintf(query,
+ unclassified_dataset[i,1],
+ unclassified_dataset[i,2],
+ unclassified_dataset[i,3],
+ unclassified_dataset[i,4],
+ unclassified_dataset[i,5])
+ dbSendQuery(con, q)
+ }
With this, we have successfully built a machine learning model within R, enriched data from CrateDB using this model, and written our enriched results back into CrateDB.